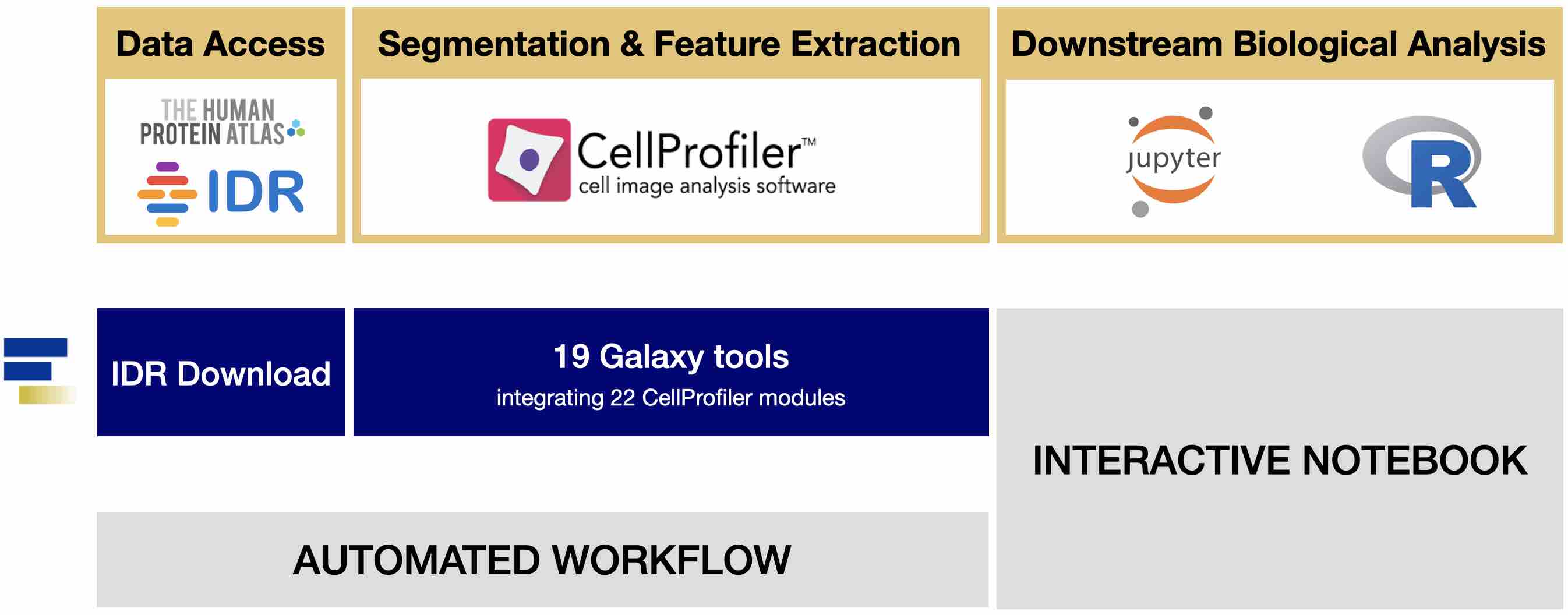
 
It is flexible enough to support running large analysis projects across multiple nodes in the Nucleus cluster, to speed up time to completion. 
#Cellprofiler 2.2.0 how toExamples of software are compilers (C, C++, F90) and software for electronic structure calculations, GAUSSIAN and GAMESS-US.įor information on how to use and write MPI and OpenMP parallel programs on UPPMAX clusters, see the tutorial.įor support, please email. CellProfiler has a batch processing feature that can be used to run jobs on a cluster independent of the graphical interface. #Cellprofiler 2.2.0 licenseThe software is mainly made available on UPPMAX computer systems, but also on other computer systems within Uppsala University, when license agreements allows it. This wastes 12 of the 112 cores available, but that is a small percentage and. We include the CellProfiler pipelines in our github repository. Its easiest to use equal size batches for processing, so we will run 100 batches of 10 image sets each, to complete our 1000 image set project. We built a CellProfiler image analysis and illumination correction pipeline (version 2.2.0) pipeline to extract these image-based features (McQuin et al., 2018). UPPMAX buys, supports and updates software for scientific computing of general interest for researchers. CellProfiler is best run using a physical core for each process, so we have 4 28 112 cores available across 4 nodes. Notice: Before you can load any bioinformatics software, you will have to issue the command "module load bioinfo-tools". More information about UPPMAX module system. List available modules with "module avail". Depending on which cluster you're logged in to you will have access to different modules/software packages. These are loaded with the " module load DESIRED_MODULE/ VERSION" command. Below you will find a list of software that's available on our resources via our module system. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
